Estimate
Base class for all estimate types, which store fit information returned from data analysis. |
|
A namedtuple container for storing a single estimate name, value, and standard deviation. |
|
A namedtuple container for storing a 1-D array of values and standard deviations for a particular parameter name. |
|
An iterable container of |
|
Contains a name and indices for a single axis of an |
|
Stores a multi-dimensional array of estimate values (and corresponding standard deviations in a separate array) for a specified parameter name. |
|
Stores a joint estimate of a collection of system parameters. |
|
Stores estimates for the confusion matrices associated with readout errors. |
|
Organizes |
|
Represents a Pauli probability distribution for some subset of qubits of a cycle of interest, as measured by KNR; see also |
|
Organizes KNR estimates into a table with columns grouped by cycle (and possibly other keywords, see |
Estimate (Parent Class)
- class trueq.estimate.base.EstimateTuple(name, val, std)
A namedtuple container for storing a single estimate name, value, and standard deviation.
- Parameters:
name (
str) – The name of the estimate.val (
float|complex) – The value of the estimate.std (
float) – The standard deviation of the estimate.
- class trueq.estimate.base.Estimate(key, options=None)
Base class for all estimate types, which store fit information returned from data analysis.
This class is not expected to be used directly, and fitting results will return subclasses such as
NormalEstimateorRCalEstimate.- Parameters:
- to_dict()
Converts the estimate to a dictionary representation.
Note
The stored
Keyis also converted to a dictionary.- Return type:
dict
- static from_dict(dic)
Converts a dictionary representation into an estimate instance.
Note
See
to_dict()for more details.
- property options
The fit options that were used to create this estimate.
- Type:
dict
Estimate Collection
- class trueq.estimate.base.EstimatesTuple(name, vals, stds, sweeps)
A namedtuple container for storing a 1-D array of values and standard deviations for a particular parameter name.
- Parameters:
name (
str) – The name of the estimates.vals (
np.ndarray) – An array of estimates.stds (
np.ndarray) – The corresponding standard deviations of the estimates.sweeps (
dict) – A dictionary mapping keyword names to lists of keyword values, where each list has the same length asvals.
- class trueq.estimate.EstimateCollection(estimates=None)
An iterable container of
Estimates with several convenience functions which make it easier to select specific estimates.import trueq as tq circuits = tq.make_srb([[0], [1, 2]], [4, 32]) circuits += tq.make_xrb([[0], [1, 2]], [4, 32]) tq.Simulator().add_overrotation(0.04).run(circuits) estimate_collection = circuits.fit() estimate_collectionTrue-Q formatting will not be loaded without trusting this notebook or rerunning the affected cells. Notebooks can be marked as trusted by clicking "File -> Trust Notebook".
Cliffords (0,)
- Key:
- labels: (0,)
- protocol: SRB
- twirl: Cliffords on [0, (1, 2)]
Cliffords (1, 2)
- Key:
- labels: (1, 2)
- protocol: SRB
- twirl: Cliffords on [0, (1, 2)]
2.8e-03 (6.3e-04) 0.0027926928633786596, 0.00062856482760666310.0e+00 (0.0e+00) 0.0, 0.01.0e+00 (8.4e-04) 0.9962764095154951, 0.0008380864368088841.0e+00 (0.0e+00) 1.0, 0.01.0e+00 (8.3e-03) 0.9960867502149159, 0.0082705851883835091.0e+00 (0.0e+00) 1.0, 0.0Cliffords (0,)
- Key:
- labels: (0,)
- protocol: XRB
- twirl: Cliffords on [0, (1, 2)]
Cliffords (1, 2)
- Key:
- labels: (1, 2)
- protocol: XRB
- twirl: Cliffords on [0, (1, 2)]
2.9e-03 (6.9e-04) 0.0028947312874968156, 0.0006926931784813809-5.8e-05 (9.7e-05) -5.776874781193708e-05, 9.652083463559954e-05-1.0e-04 (2.9e-04) -0.000102038424118156, 0.00029108434690042025.8e-05 (9.7e-05) 5.776874781193708e-05, 9.652083463559954e-051.0e+00 (7.8e-04) 1.0002721163467685, 0.00077630412983590241.0e+00 (2.1e-04) 0.9998767635643779, 0.000205899218662069081.0e+00 (1.1e-02) 1.0119035653771058, 0.0108305919083257951.1e+00 (4.7e-03) 1.0613191254240442, 0.004692647687300659
- Parameters:
estimates (
NoneType|Estimate-like |Iterable) – Either a singleEstimateor an iterable of several estimates. IfNoneis provided, theEstimateCollectionwill be empty.
- append(estimate)
Appends a single
Estimateor an iterable of several estimates to the collection.
- keys(**filter)
Returns the set of all keys of this estimate collection matching the given filter.
import trueq as tq circuits = tq.make_srb([[0], [1, 2]], [4, 32]) circuits += tq.make_xrb([[0], [1, 2]], [4, 32]) tq.Simulator().add_overrotation(0.04).run(circuits) estimate_collection = circuits.fit() estimate_collection.keys(protocol="SRB")True-Q formatting will not be loaded without trusting this notebook or rerunning the affected cells. Notebooks can be marked as trusted by clicking "File -> Trust Notebook".
protocol The characterization protocol used to generate a circuit.labels twirl The twirling group used to generate a circuit.
- Key:
- labels: (0,)
- protocol: SRB
- twirl: Cliffords on [0, (1, 2)]
SRB (0,) Cliffords on [0, (1, 2)]
- Key:
- labels: (1, 2)
- protocol: SRB
- twirl: Cliffords on [0, (1, 2)]
SRB (1, 2) Cliffords on [0, (1, 2)]
- Parameters:
**filter – The filter on keys.
- Return type:
- subset(name_pattern=None, filter_fn=None, **filter)
Returns an
EstimateCollectioncontaining only the estimates in this collection that match the filter on keys, and whose parameter names match the given filter function or the pattern.
- Parameters:
name_pattern (
string) – A regular expression that specifies a pattern for the names, for example"(p_ZZ.*)|(e_F)"to match any name that starts withp_ZZor any name equal toe_F.filter_fn (
function) – A function which accepts anEstimateand returns eitherTrueorFalse.**filter – The filter on keys.
- Return type:
- one_or_none(filter_fn=None, **filter)
Returns a single estimate if it is the only estimate in this collection that matches the filter. Otherwise returns
None.
- sorted(*names)
Returns an
EstimateCollectionsorted by the keys contained in the estimates, seetrueq.KeySet.sorted()for more information.
- Parameters:
*names – One or more name strings that should take priority in sorting.
- Return type:
- update_keys(*other, keep=None, remove=None, **kwargs)
Updates every estimate’s
keyin this collection with new keywords/values. If a given key does not have a given keyword, it is added. If it already exists, it is overwritten. See alsotrueq.Key.copy()which this method uses.import trueq as tq # generate an estimate collection by calling fit() on some circuits circuits = tq.make_srb([[0], [1, 2]], [4, 32]) circuits += tq.make_xrb([[0], [1, 2]], [4, 32]) tq.Simulator().add_overrotation(0.04).run(circuits) estimates = circuits.fit() # give each circuit a new keyword 'banana' with value 10 estimates.update_keys(banana=10) estimates.keys()True-Q formatting will not be loaded without trusting this notebook or rerunning the affected cells. Notebooks can be marked as trusted by clicking "File -> Trust Notebook".
protocol The characterization protocol used to generate a circuit.labels twirl The twirling group used to generate a circuit.banana
- Key:
- banana: 10
- labels: (0,)
- protocol: SRB
- twirl: Cliffords on [0, (1, 2)]
SRB (0,) Cliffords on [0, (1, 2)] 10
- Key:
- banana: 10
- labels: (1, 2)
- protocol: SRB
- twirl: Cliffords on [0, (1, 2)]
SRB (1, 2) Cliffords on [0, (1, 2)] 10
- Key:
- banana: 10
- labels: (0,)
- protocol: XRB
- twirl: Cliffords on [0, (1, 2)]
XRB (0,) Cliffords on [0, (1, 2)] 10
- Key:
- banana: 10
- labels: (1, 2)
- protocol: XRB
- twirl: Cliffords on [0, (1, 2)]
XRB (1, 2) Cliffords on [0, (1, 2)] 10
- Parameters:
*other – One or more dict-like objects (e.g.
Keyordict) to update the keys with. Updating is applied in the given order. If a name specified in any of these objects already exists after thekeeporremoveprocess has taken place, it is updated.keep (
str|list) – A string or list of strings specifying the only names to keep during the updates. By default, all names are kept. Only one of the optionskeeporremovemay be used.remove (
str|list) – A string or list of strings specifying names to remove during the updates. By default, no names are removed. Only one of the optionskeeporremovemay be used.**kwargs – Name-value items to update the keys with. If a name specified here already exists after the
keeporremoveprocess has taken place, it is updated.- Returns:
This estimate collection.
- Return type:
- Raises:
ValueError – If the mutally exclusive
keepandremoveare both set toTrue.
- array(name, *axes, name_axis=0)
Constructs a multi-dimensional array of estimate values for a specified parameter name. Each axis corresponds to a sweep over metadata values from a particular
Keyname from the estimates in this collection, or over parameter names themselves ifnameis provided as a pattern with more than one match.import trueq as tq circuits = tq.make_srb([0, 1, 2, 3, 4], [4, 32]) circuits += tq.make_xrb([0, 1, 2, 3, 4], [4, 32]) tq.Simulator().add_overrotation(0.04).run(circuits) fit = circuits.fit() # extract a 2-d array of 'p' estimates with first axis over qubit labels, # and second axis over protocol arr = fit.array("p", "labels", "protocol") # we can look at the axes and value array directly print(arr.axes) print(arr.vals, "\n") # we can also look at the html representation, which condenses all but the # last axis into rows, and displays the last axis as columns arr(ArrayAxis('labels', ((0,), (1,), (2,), (3,), (4,))), ArrayAxis('protocol', ('SRB',))) [[0.99721427] [0.99488382] [0.99615792] [0.99744018] [0.99570605]]True-Q formatting will not be loaded without trusting this notebook or rerunning the affected cells. Notebooks can be marked as trusted by clicking "File -> Trust Notebook".
protocol SRB labels (0,) 9.97e-01 (9.88e-04) val=0.997214
std=0.00098836(1,) 9.95e-01 (1.34e-03) val=0.994884
std=0.00134008(2,) 9.96e-01 (9.24e-04) val=0.996158
std=0.000923794(3,) 9.97e-01 (1.16e-03) val=0.99744
std=0.00116156(4,) 9.96e-01 (1.12e-03) val=0.995706
std=0.00112492Note
The extracted array is always (hyper-)rectangular, whereas estimate collections are often ragged. When a particular value does not exist in the collection,
np.nanis inserted as the array element.
- Parameters:
name (
str|ArrayAxis) – The name of the parameter to extract, e.g."e_F", a regular expression whose matched named are extracted, e.g."e_F|p"or"e__[IXYZ]+", or anArrayAxiswhose values are parameter names.*axes – A sequence of strings or
ArrayAxisthat are used to index axes of the array. Each axis (except possibly one defined byname, which is a special case) corresponds to metadata values from a specific keyword found in thekeys()of this collection. For example, a string axis value of"protocol"will cause the corresponding axis to be over different protocols sorted by protocol name, or a string axis value of"labels"will be over sorted subsystem labels. For more fine-grained control, axes can be manually specified byArrayAxisinstances.name_axis (
int) – In the case where multple names are matched, either becausenameis a regular expression with multiple matching names or becausenamewas given as anArrayAxiswith multiple values, which axis to put the names on. The default is the 0’th axis, and the remaining axes are ordered according to the order of theaxes.- Return type:
- Raises:
ValueError – If no estimates with names matching
nameare found.ValueError – If multiple estimates with the same name are found that do not differ on their axis values.
- to_dict_list()
Returns a list of dictionary representations of the
Estimateobjects in this collection.
- Return type:
list
- static from_dict_list(dict_list)
Creates a new estimate collection from a list of dictionary representations of
Estimateobjects.
- Parameters:
dict_list (
list) – A list of dictionary representations of estimate objects.- Return type:
- property plot
An object that stores all plotting functions deemed relevant to this
EstimateCollection. If one of these functions is called, the data from this estimate collection is analyzed and used.
- Type:
Estimate Array
- class trueq.estimate.ArrayAxis(name, indices, is_name_axis=False)
Contains a name and indices for a single axis of an
EstimateArray.- Parameters:
name (
str) – The name of the axis.indices (
Iterable) – The indexing values of the axis. These can be of any type. The length should match the dimension of the axis.is_name_axis (
bool) – Whether this axis is over parameter names; at most one such axis can exist for anEstimateArray.
- property name
The name of this axis.
- Type:
str
- property size
The size of this axis.
- Type:
int
- property indices
The indexing values of this axis.
- Type:
tuple
- property is_name_axis
Whether this axis is over parameter names.
bool
- class trueq.estimate.EstimateArray(vals, stds, axes)
Stores a multi-dimensional array of estimate values (and corresponding standard deviations in a separate array) for a specified parameter name. Each axis corresponds to a sweep over metadata values from a particular
Keyname from the estimates in anEstimateCollection, or over parameter names themselves. Instances of this class are typically constructed byarray().- Parameters:
vals (
array_like) – An array of parameter values estimates.stds (
array_like) – The standard deviations of each member ofvals, an array with the same shape asvals.axes (
Iterable) – A sequence ofArrayAxisinstances whose sizes must match the shape ofvalsandstds.
- property stds
An array of parameter value standard deviations, with the same shape as
vals. The axes are described byaxes.- Type:
array_like
- property vals
An array of parameter values, with the same shape as
vals. The axes are described byaxes.- Type:
array_like
- nan_reason(idxs)
Returns a reason for why the array element at the given index is
numpy.nan. If it is a number, then an empty string is returned.- Parameters:
idxs (
tuple) – The nd-index to address.- Return type:
str
- to_dataframe(include_vals=True, include_stds=True)
Converts this array to a dataframe.
If this array has two or more dimensions, the last dimension is used for the columns of the dataframe, and the first dimensions are flattened together to form the rows, and indexed with a
pandas.MultiIndex. If this array has only one dimension, then the values are put into rows.If both
include_valsandinclude_stdsareTrue, which is the default, then they appear in separate columns, where apandas.MultiIndexis used along with the last dimension of this array if it has at least two dimensions in total.- Parameters:
include_vals (
bool) – Whether to include estimate values in the dataframe.include_stds (
bool) – Whether to include standard deviations in the dataframe.
- Return type:
pandas.DataFrame
Normal Estimates
- class trueq.estimate.NormalEstimate(key, names, values, err=None, raw=None, options=None)
Stores a joint estimate of a collection of system parameters. Each parameter is described by a name and an estimated mean value. These mean values are assumed to be normally distributed, and their covariance matrix (or vector of standard deviations for uncorrelated estimates) is optionally present.
import trueq as tq estimate = tq.NormalEstimate( tq.Key(), ["a", "b", "c"], [1, 2, 3], [0.2, 0.2, 0.2] ) # get the estimated value of b estimate.b
EstimateTuple(name='b', val=2, std=0.2)
- Parameters:
key (
Key) – A key used for record keeping in analysis. For example,Key(protocol='SRB', n_random_cycles=10).names (
Iterable) – An iterable collection of strings which describe the parameters returned from fitting.values (
list|numpy.ndarray) – An iterable containing the estimated values associated with the provided names.err (
NoneType|numpy.ndarray|list) – Optional array or list containing either the covariance matrix or the standard deviation vector associated with the estimated values. IfNoneis provided then this is set to a vector of zeros.raw (
NoneType|dict) – Optional, the data used to estimate the values. If not provided, this defaults to an empty dictionary.options (
NoneType|dict) – Options that were passed tofit()method.
- Raises:
ValueError – If the lengths of
namesanderr(if provided) do not match the length ofvalues.
- property names
An iterable collection of strings which describe the parameters returned from fitting.
- Type:
Iterable
- property values
An iterable containing the estimated values associated with the provided names.
- Type:
Iterable
- property raw
The data used to estimate the values. The format of this dictionary may vary depending on who constructed it. Some examples of the format used by True-Q™ protocols are:
Protocols
Format
{<sequence_length>: [<circuit expectation values>], ...}{<measurement_basis>: {<sequence_length>: [<circuit expectation values>], ...}, ...}- Type:
dict
- property err
Array containing either the covariance matrix or the standard deviation vector associated with the estimated values. If
err=Nonewas provided, this will be a vector of zeros.- Type:
numpy.ndarray
- property cov
The covariance matrix of this estimate.
If
err=Nonewas provided, this will be an array of zeros.- Type:
numpy.ndarray
- property std
A vector of standard deviations for every element of this estimate.
If
err=Nonewas provided, this will be a vector of zeros.If a covariance matrix is present then this is the square root of diagonal terms.
- Type:
numpy.ndarray
- subset(name_pattern)
Returns a new
NormalEstimatethat contains only the parameters whose name matchesname_pattern.- Parameters:
name_pattern (
str) – A regular expression that specifies a pattern for the names.- Type:
py:class:~trueq.estimate.NormalEstimate
Readout Estimates
- class trueq.estimate.RCalEstimate(key, values, options=None)
Stores estimates for the confusion matrices associated with readout errors.
- Parameters:
key (
Key) – A hashable object used for record keeping in analysis. For example,Key(protocol='SRB', n_random_cycles=10).values (
dict) – A dict where the keys of the dict are the single qubit labels, and the values are the associated confusion matrices for the probabilities of correctly labeling the qubit’s final state.options (
NoneType|dict) – Options that were passed tofit()method.
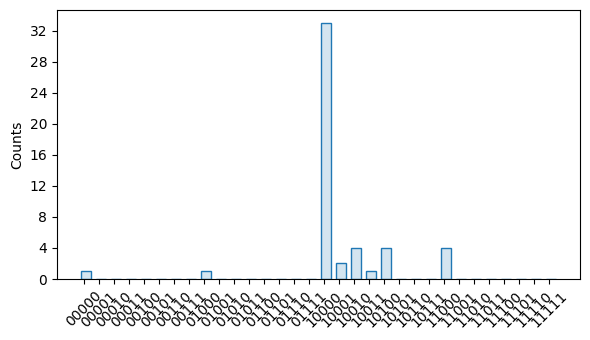
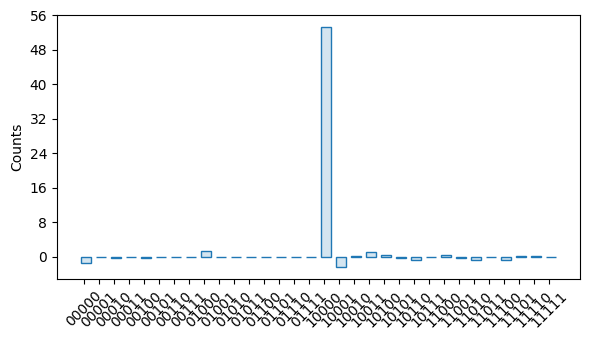
- apply_correction(results, labels=None)
Applies this calibration as a correction to the given results. This is done by inverting each calibration matrix and contracting it onto the corresponding indices of the given results.
import trueq as tq # make a test and RCAL circuits rcal_circuits = tq.make_rcal(range(5)) circuit = tq.Circuit([{0: tq.Gate.x}, {range(5): tq.Meas()}]) # construct a simulator with readout error and run the circuits sim = tq.Simulator().add_readout_error([0.1, 0.05]) sim.run(rcal_circuits, 10000) sim.run(circuit) # create an RCAL estimate rcal_est = rcal_circuits.fit()[0] # plot original and corrected bitstring distributions circuit.results.plot() rcal_est.apply_correction(circuit.results).plot()
Note
This method necessarily has exponential scaling in the number of qubits. Further, no clipping is performed, so that the output distribution may contain small negative values due to finite sampling effects.
- Parameters:
results (
Results) – A results object to correct.labels (
Iterable|NoneType) – An optional labelling (iterable of unique non-negative integers) of the bitstrings in the given results object. These labels are used to decide which correction matrices to apply where. IfNone, the labels from the key of this estimate are assumed.
- Returns:
A new results object equal to the correction of the input.
- Return type:
- Raises:
ValueError – If no
labelsare given and there are no labels in the key of this instance.ValueError – If bitstring lengths are not compatible with the number of labels.
- property values
A dict where the keys of the dict are the single qubit labels, and the values are the associated confusion matrices for the probabilities of correctly labeling the qubit’s final state.
- Type:
dict
Comparison Table
- class trueq.estimate.comp_table.CompTable(fit_or_circuits)
Organizes
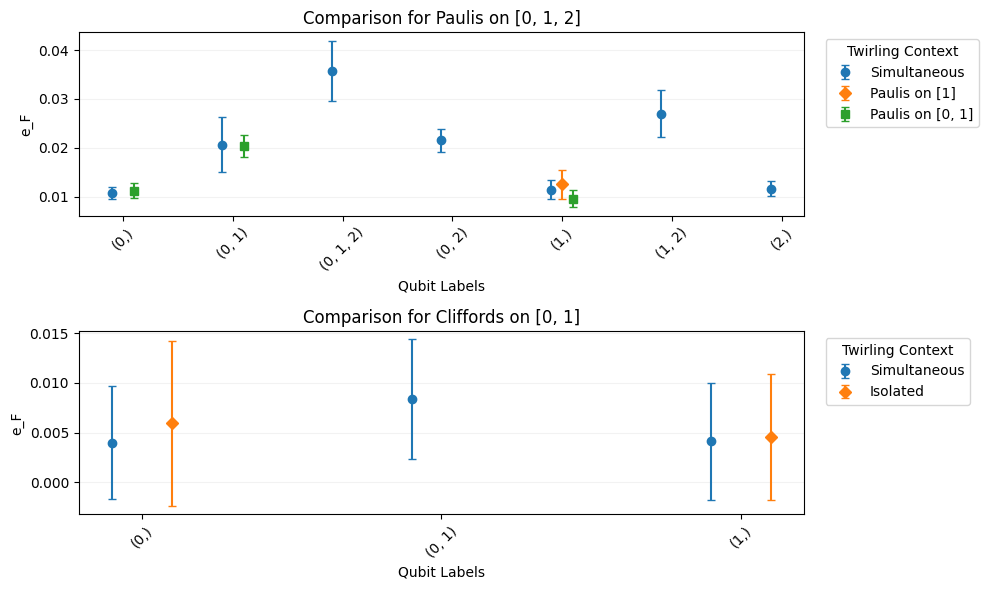
NormalEstimates into a parent-children structure based on twirling groups in a way that makes it easier to compare values.from itertools import chain, combinations import trueq as tq # generate CB circuits circuits = tq.make_cb({range(3): tq.Gate.x}, [4, 32]) circuits += tq.make_cb({range(2): tq.Gate.x}, [4, 32]) circuits += tq.make_cb({1: tq.Gate.x}, [4, 32]) # use a different twirling group circuits += tq.make_cb({range(2): tq.Gate.x}, [4, 32], twirl="C") circuits += tq.make_cb({0: tq.Gate.x}, [4, 32], twirl="C") circuits += tq.make_cb({1: tq.Gate.x}, [4, 32], twirl="C") # add a different qubit circuits += tq.make_cb({range(3): tq.Gate.x}, [4, 32]) # run the circuits on a noisy simulator tq.Simulator().add_overrotation(0.04).run(circuits) def powerset(iterable): # powerset([1,2,3]) -> () (1,) (2,) (3,) (1, 2) (1, 3) (2, 3) (1, 2, 3) s = list(iterable) return chain.from_iterable(combinations(s, r) for r in range(len(s) + 1)) # get infidelity on every possible combination of labels fit = circuits.fit(set(chain(powerset(range(3)), powerset((0, 5, 2))))) # this plotter visualizes CompTable; every subplot is a Family, and every trace # is a Member fit.plot.compare_twirl("e_F")
- Parameters:
fit_or_circuits (
CircuitCollection|EstimateCollection) – An estimate collection or set of circuits to create the data table for.
- class Family(parent, title, labels, names, members, protocols)
- labels
Alias for field number 2
- members
Alias for field number 4
- names
Alias for field number 3
- parent
Alias for field number 0
- protocols
Alias for field number 5
- title
Alias for field number 1
- class Member(title, twirls, data, protocol)
- data
Alias for field number 2
- protocol
Alias for field number 3
- title
Alias for field number 0
- twirls
Alias for field number 1
- table(names=None)
Returns all data in a structure.
import trueq as tq circuits = tq.make_crosstalk_diagnostics([0, 1, 2], [4, 32]) tq.Simulator().add_overrotation(0.05).run(circuits) data = tq.estimate.comp_table.CompTable(circuits) for family in data.table(["p", "r"]): print(family.title) print(family.labels) for member in family.members: print(member.title) print(member.data) print("")
Cliffords on [0, 1, 2] [(0,), (1,), (2,)] Simultaneous {'p': {(0,): EstimateTuple(name='p', val=0.9939236346479658, std=0.0015423185968745058), (1,): EstimateTuple(name='p', val=0.9931414377622905, std=0.0015458536528304013), (2,): EstimateTuple(name='p', val=0.9936729978672539, std=0.0011465811395867723)}} Isolated {'p': {(0,): EstimateTuple(name='p', val=0.9939023519241923, std=0.0013221758030465519), (1,): EstimateTuple(name='p', val=0.9961954897978763, std=0.001114238798559842), (2,): EstimateTuple(name='p', val=0.9937304705075709, std=0.0015425107107473216)}}- Parameters:
names (
str|Iterable|None) – An estimate parameter of names or list thereof to include in the output. All names are included by default.- Return type:
list
KNR Specializations
- class trueq.estimate.knr.KnrBodyEstimate(key, names, values, err=None, raw=None, options=None)
Represents a Pauli probability distribution for some subset of qubits of a cycle of interest, as measured by KNR; see also
make_knr(). Thekeystores all relevant metadata, including:key.cycleis the cycle of interest.key.twirlis the twirling group, which may include a superset of the qubits referenced by the cycle.key.labelsare the sorted qubit labels defining the subsystem for which this estimate is a distribution over.key.subsystemsare the combinations of labels for which marginal distributions are constructed.
- property subcycle
A subcycle of this estimate’s cycle of interest containing only the gate(s) that this estimate describes. Any idling qubits (i.e. qubits which are twirled by the protocol but absent from the cycle of interest) that this estimate describes are added as single qubit identity gates.
- Type:
- class trueq.estimate.knr.KnrDataTable(fit_or_circuits, group_by=('name', 'cycles', 'twirl'))
Organizes KNR estimates into a table with columns grouped by cycle (and possibly other keywords, see
group_by) and rows grouped by error modes.- Parameters:
fit_or_circuits (
CircuitCollection|EstimateCollection) – An estimate collection or set of circuits to create the data table for.group_by (
Iterable) – Which keywords to group columns by. For example, if the estimate collection contains data for the same cycle for multiple chips, and this information is distinguished by a “chip” keyword, then this value could be (“cycles”, “chip”, “twirl”). The keywords “cycles” and “twirl” should appear in this list.
- class Cell(mean, std, subcycles)
Storage type for the contents of a single cell in the table
- mean
Alias for field number 0
- std
Alias for field number 1
- subcycles
Alias for field number 2
- class Row(sort_key, degens, param_names, latex)
Storage type for row descriptions, which index Cell rows.
- degens
Alias for field number 1
- latex
Alias for field number 3
- param_names
Alias for field number 2
- sort_key
Alias for field number 0
- set_truncation(cutoff, relative_to_max=True, cl=None)
Sets the cutoff at which rows are hidden. If every probability in a subrow falls below the cutoff across an entire row of cells, then these subrows are removed when
get_cell()androw_infoare called.- Parameters:
cutoff (
float) – The cutoff value to use.relative_to_max (
bool) – Whether the cutoff is relative to the maximum probability in the whole table (in which case the cutoff used iscutoff * max_val). Otherwise, the cutoff is absolute.cl (
float|NoneType) – The confidence level at which to cutoff, orNoneto use the current default accessed byget_cl().
- property n_cell_rows
The number of cell rows in this table.
Note
This value may change after
set_truncation()is called.- Type:
int
- property n_cell_cols
The number of cell columns in this table.
- Type:
int
- property row_info
A list of
Rowinstances which describe the contents of each cell row of the table.Note
This value may change after
set_truncation()is called.- Type:
list
- get_cell(idx_cell_row, idx_cell_col)
Gets a single cell by index. This cell contains a matrix of probabilities, a matrix of their standard deviations, and label information for each column in the matrix. Note that row information is found in the corresponding element of
row_info, and a header for the whole cell is found in the corresponding element ofcol_info.Note
Cell row indices may change when
set_truncation()is called.- Parameters:
idx_cell_row (
int) – An index belown_cell_rows.idx_cell_col (
int) – An index belown_cell_cols.
- Return type:
- property col_info
A list of
Colinstances which describe the contents of each cell row of the table.- Type:
list
- row_max(idx_cell_row)
Returns the maximum error probability of the given row and the corresponding standard deviation.
- Parameters:
idx_cell_row (
int) – An index belown_cell_rows.- Return type:
tuple
- property max
The maximum error probability in the entire table and its corresponding standard deviation.
- Type:
float
- property heights
A list of heights of each cell row.
- Type:
list
- property widths
A list of the maximum widths of each cell column.
- Type:
list
